prepared by Svetlana Ugarcina Perovic
The AMR March digest features the latest publications on global health risk of antibiotic resistance genes, bacterial responses to antibiotic combinations, Hi-C assisted metagenomics for AMR resistance tracking and more. Happy #AMR reading!
Join us for the spring #EMBARK2022 webinars!
Global
Assessment of global health risk of antibiotic resistance genes – Zhenyan Zhang – Nature Communications
*Authors developed a novel method for quantitatively surveilling the health risk of ARGs, by integrating human accessibility, mobility, pathogenicity and clinical availability. Antibiotic resistance risk was detected all around the world, even in the polar region. Human-associated habitats posed the highest risk of antibiotic resistance than other habitats.
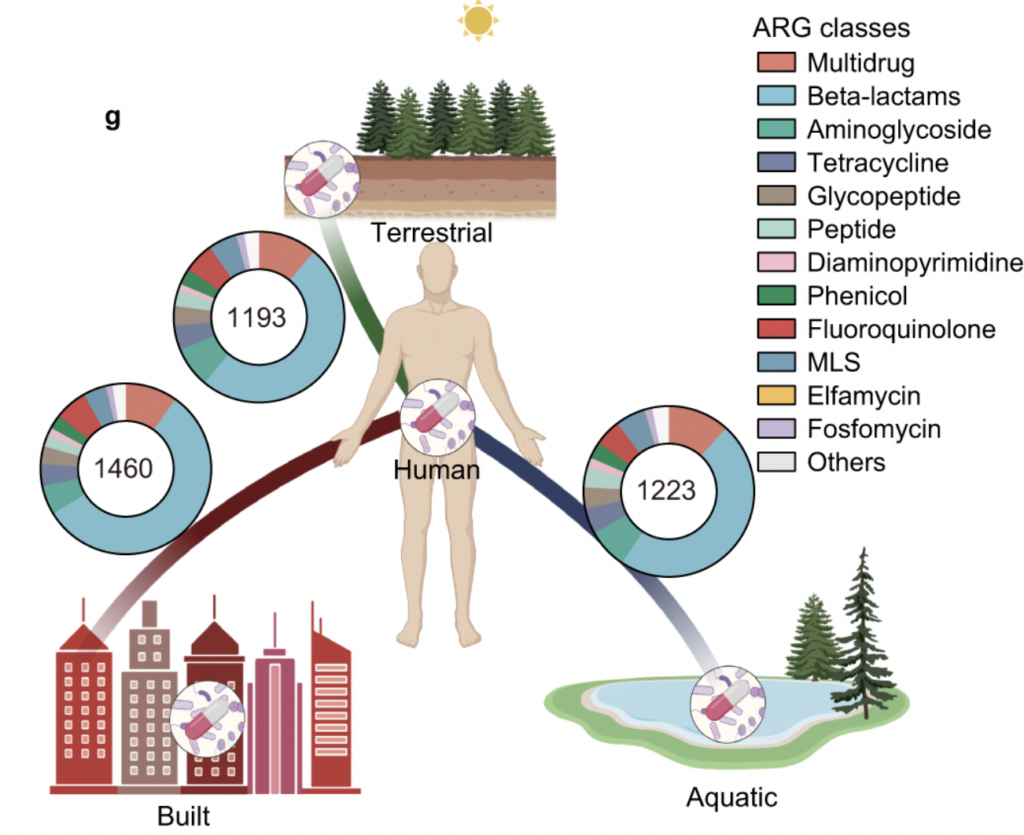
Number in the circles represents the number of shared ARGs.
Existing evidence on antibiotic resistance exposure and transmission to humans from the environment: a systematic map – Isobel Catherine Stanton – Environmental Evidence
Landscape of mobile genetic elements and their antibiotic resistance cargo in prokaryotic genomes – Supriya Khedkar, Georgy Smyshlyaev, Ivica Letunic, Oleksandr M Maistrenko, Luis Pedro Coelho, Askarbek Orakov, Sofia K Forslund, Falk Hildebrand, Mechthild Luetge, Thomas S B Schmidt, Orsolya Barabas, Peer Bork – Nucleic Acids Research
Intra- and interpopulation transposition of mobile genetic elements driven by antibiotic selection – Yi Yao – Nature Ecology & Evolution
Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM – Mislav Acman – Nature Communications
Clinical
Phenotypic and Genotypic Characterization of a Hypervirulent Carbapenem-Resistant Klebsiella pneumoniae ST17-KL38 Clinical Isolate Harboring the Carbapenemase IMP-4 – Jintao He – Antimicrobial Chemotherapy
The repurposing of Tebipenem pivoxil as alternative therapy for severe gastrointestinal infections caused by extensively drug resistant Shigella spp. – Elena Fernández Alvaro – eLife
Urinary culture sensitivity after a single empirical antibiotic dose for upper or febrile urinary tract infection: a prospective multicenter observational study – John Gregor – Clinical Microbiology and Infection
Review: The physiology and genetics of bacterial responses to antibiotic combinations – Roderich Roemhild – Nature Reviews Microbiology
*Authors discuss the effects of antibiotic combinations, previous treatments and the role of stress-response systems as well as resistance mutation–drug interactions.
Decreased efficacy of antimicrobial agents in a polymicrobial environment – Thomas James O’Brien – The ISME Journal
Human microbiome
Strain-level characterization of broad host range mobile genetic elements transferring antibiotic resistance from the human microbiome – Samuel C. Forster – Nature Communications
Review: Good microbes, bad genes? The dissemination of antimicrobial resistance in the human microbiome – Alexander Crits-Christoph – Gut Microbe
Inflammatory Endotype-Associated Airway Resistome in Chronic Obstructive Pulmonary Disease – Xinzhu Yi – Microbiology Spectrum
Animal
Extensive metagenomic analysis of the porcine gut resistome to identify indicators reflecting antimicrobial resistance – Yunyan Zhou – Microbiome
Novel canine high-quality metagenome-assembled genomes, prophages and host-associated plasmids provided by long-read metagenomics together with Hi-C proximity ligation – Anna Cuscó – Microbial Genomics
Animal experiments
Antibiotics cause metabolic changes in mice primarily through microbiome modulation rather than behavioral changes – Kale S. Bongers – PlosOne
Phages
A Phage Foundry framework to systematically develop viral countermeasures to combat antibiotic resistant bacterial pathogens – Vivek K. Mutalik and Adam P. Arkin – iScience
Water
Characteristics of Antibiotic Resistance Genes and Antibiotic-Resistant Bacteria in Full-Scale Drinking Water Treatment System Using Metagenomics and Culturing – Qihui Gu – Frontiers in Microbiology
Municipal Wastewaters Carry Important Carbapenemase Genes Independent of Hospital Input and Can Mirror Clinical Resistance Patterns – Adela Teban-Man, Edina Szekeres, Peiju Fang, Uli Klümper, Adriana Hegedus, Andreea Baricz, Thomas Ulrich Berendonk, Marcel Pârvu, Cristian Coman – Environmental Microbiology
Soil
Cross-biome antibiotic resistance decays after millions of years of soil development – Qing-Lin Chen – The ISME Journal
Metagenomic strategies identify diverse integron-integrase and antibiotic resistance genes in the Antarctic environment – Verónica Antelo – MicrobiologyOpen
Techniques
HAM-ART: An optimised culture-free Hi-C metagenomics pipeline for tracking antimicrobial resistance genes in complex microbial communities – Lajos Kalmar – Plos Genetics
*A comprehensive laboratory and bioinformatics pipeline HAM-ART (Hi-C Assisted Metagenomics for Antimicrobial Resistance Tracking) optimised for the generation of metagenome-assembled genomes including both chromosomal and extrachromosomal AMR genes. Authors found significant differences in the distribution of AMR genes between low- and high-antimicrobial use farms including a plasmid-borne lincosamide resistance gene exclusive to high-antimicrobial use farms in three species of Lactobacilli.
Comparing Long-Read Assemblers to Explore the Potential of a Sustainable Low-Cost, Low-Infrastructure Approach to Sequence Antimicrobial Resistant Bacteria With Oxford Nanopore Sequencing – Ian Boostrom – Frontiers in Microbiology
Bioinformatics
IntegronFinder 2.0: identification and analysis of integrons across Bacteria, with a focus on antibiotic resistance in Klebsiella – Bertrand Néron – bioRxiv
Whole genome sequencing and gene sharing network analysis powered by machine learning identifies antibiotic resistance sharing between animals, humans and environment in livestock farming – Zixin Peng – Plos Computational Biology
*Authors developed a computational framework identifying known AMR genes as well as yet unknown genetic determinants underlying AMR in E. coli from complex anthropogenic environments.
Events
Recordings of the EDAR6 count down webinars available here.
Antimicrobial Resistance – Genomes, Big Data and Emerging Technologies (Virtual Conference): 27–29 April 2022, Wellcome Genome Campus, UK
Podcast