AMR MARCH, 2023
prepared by Ulrike Löber
March digest features a review of novel methods to quantifying the physical linkage between specific antimicrobial resistance genes and mobile genetic elements, how traveling affects AMR, and more.
We are looking for speakers for our EMBARK webinars in 2023. If you want to share your groundbreaking research on antimicrobial resistance, please get in touch with us!
General
Latent antibiotic resistance genes are abundant, diverse, and mobile in human, animal, and environmental microbiomes – Juan Salvador Inda-Díaz, Marcos Parras-Moltó, Anna Johnning, Johan Bengtsson-Palme & Erik Kristiansson – Microbiome
Prevalence of Plasmid-Associated Tetracycline Resistance Genes in Multidrug-Resistant Escherichia coli Strains Isolated from Environmental, Animal and Human Samples in Panama – I. E. Ramírez-Bayard – Antibiotics
Perspective: Challenges in Forecasting Antimicrobial Resistance – Sen Pei – Emerging Infectious Diseases
Determination and quantification of microbial communities and antimicrobial resistance on food through host DNA-depleted metagenomics – Samuel J. Bloomfield – Food Microbiology
Editorial: Antimicrobial resistance and antimicrobial alternatives – Marwan Osman – Frontiers in Medicine
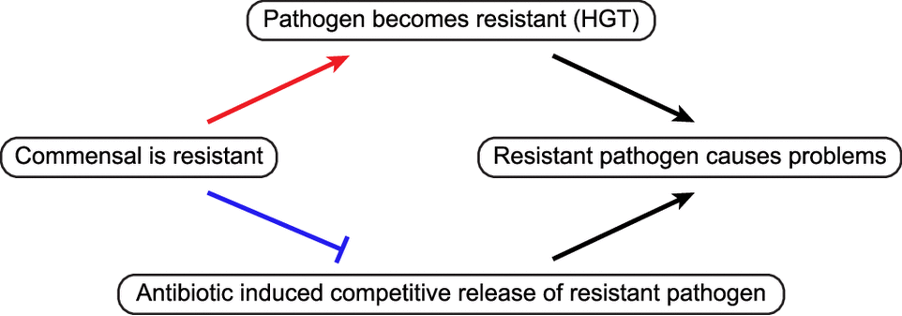
Review: Reframing antimicrobial resistance as a continuous spectrum of manifestations – Sarah M Schrader – Current Opinion in Microbiology
Intermittent antibiotic treatment of bacterial biofilms favors the rapid evolution of resistance – Masaru Usui – Communications Biology
Heavy Metal Pollution Impacts Soil Bacterial Community Structure and Antimicrobial Resistance at the Birmingham 35th Avenue Superfund Site – Anuradha Goswami – Microbiology Spectrum
Horizontal gene transfer
Review: How do interactions between mobile genetic elements affect horizontal gene transfer? – Tanya Horne – Current Opinion in Microbiology
Antimicrobial peptides
Development of novel broad-spectrum antimicrobial lipopeptides derived from plantaricin NC8 β – Emanuel Wiman – Scientific Reports
Global health
*Preprint: Within-host density and duration of colonization of multidrug-resistant Enterobacterales acquired during travel to the tropics – Olivier Cotto, Laurence Armand-Lefèvre, Sophie Matheron, Etienne Ruppé, François Blanquart the VOYAG-R study group – bioRxiv
Review: Reservoirs of antimicrobial resistance in the context of One Health – Milena Despotovic – Current Opinion in Microbiology
Alterations in faecal microbiome and resistome in Chinese international travellers: a metagenomic analysis – Man Kit Cheung – Journal of Travel Medicine
Human-associated AMR
Functional metagenomic libraries generated from anthropogenically impacted environments reveal importance of metabolic genes in biocide and antibiotic resistance – Aimee K. Murray – Current Research in Microbial Sciences
Population-level impacts of antibiotic usage on the human gut microbiome – Kihyun Lee – Nature Communications
Genomic diversity of non-diarrheagenic fecal Escherichia coli from children in sub-Saharan Africa and south Asia and their relatedness to diarrheagenic E. coli – Tracy H. Hazen – Nature Communications
Integrative omics identifies conserved and pathogen-specific responses of sepsis-causing bacteria – Andre Mu – Nature Communications
Development of the oral resistome during the first decade of life – Smitha Sukumar – Nature Communications
Mutations Related to Antibiotics Resistance in Helicobacter pylori Clinical Isolates from Bangladesh – Kartika Afrida Fauzia – Antibiotics
Hospital
Extended-Spectrum β-Lactamase Genes Traverse the Escherichia coli Populations of Intensive Care Unit Patients, Staff, and Environment – Robert A. Moran – Microbiology Spectrum
Rectal carriage of extended-spectrum β-lactamase-producing Enterobacteriales among neonates admitted into a special care baby unit, southwest Nigeria – Temitope O Obadare – Transactions of The Royal Society of Tropical Medicine and Hygiene
Virulence analysis and antibiotic resistance of Klebsiella pneumoniae isolates from hospitalised patients in Poland – Barbara Kot – Scientific Reports
Animal
Plasmid-Mediated Colistin Resistance Genes mcr-1 and mcr-4 in Multidrug-Resistant Escherichia coli Strains Isolated from a Healthy Pig in Portugal – Ana Amaro – Microbial Drug Resistance
Phenotypic and genotypic characterization of antibiotic resistance of Salmonella Heidelberg in the south of Brazil – Luana Sielski Galvão Soares – International Journal of Food Microbiology
Trophic level and proteobacteria abundance drive antibiotic resistance levels in fish from coastal New England – Benjamin J. Korry – Animal Microbiome
Longitudinal study of the short- and long-term effects of hospitalisation and oral trimethoprim-sulfadiazine administration on the equine faecal microbiome and resistome – Mathijs J. P. Theelen – Microbiome
In vitro digestion of ESC-resistant Escherichia coli from poultry meat and evaluation of human health risk – May Linn Buberg – Frontiers in Microbiology
Identification and characterization of the causative agents of Focal Ulcerative Dermatitis in commercial laying hens – Diana I. Ayala – Frontiers in Veterinary Science
Water
Microbiome, resistome and mobilome of chlorine-free drinking water treatment systems – David Calderón-Franco – Water Research
Review: Sewage surveillance of antibiotic resistance holds both opportunities and challenges – D. G. Joakim Larsson – Nature Reviews Microbiology
Preprint: Antibiotic Resistance Gene Variant Sequencing is Necessary to Reveal the Complex Dynamics of Immigration from Sewers to Activated Sludge – Claire Gibson – bioRxiv
Evidence for wastewaters as environments where mobile antibiotic resistance genes emerge – Fanny Berglund – Communications Biology
Viruses and phages
The persistence and stabilization of auxiliary genes in the human skin virome – Ema H. Graham – Virology Journal
Altered infective competence of the human gut microbiome in COVID-19 – Laura de Nies – Microbiome
Review: Antimicrobial resistance in patients with COVID-19: a systematic review and meta-analysis – Bradley J Langford – The Lancet Microbe
Characterization of antibiotic resistomes by reprogrammed bacteriophage-enabled functional metagenomics in clinical strains – Gábor Apjok – Nature Microbiology
Bioinformatics
Prediction of vancomycin initial dosage using artificial intelligence models applying ensemble strategy – Wen-Hsien Ho – BMC Bioinformatics
Protocols
Nanopore Sequencing Discloses Compositional Quality of Commercial Probiotic Feed Supplements – Worarat Kruasuwan – Scientific Reports
*Quantification of the mobility potential of antibiotic resistance genes through multiplexed ddPCR linkage analysis – Magali de la Cruz Barron, David Kneis, Alan Xavier Elena, Kenyum Bagra, Thomas U Berendonk, Uli Klümper – FEMS Microbiology Ecology
News and Views
European Science-Media Hub – Interview with Prof. Francesco Imperi on new strategies to fight antimicrobial resistance
Antibiotic Resistance: A mobile target – Carolina Oliveira de Santana – eLife
Podcasts
The AMR Studio Episode 46 – Alison Prendiville & service design. Antimicrobials in food animals. Bacteriuria & ICU stays.
EDITORS IN CONVERSATION Episode 63 – Antimicrobial Resistance in Neisseria gonorrhoeae
Webinars
WHO costing & budgeting tool for national action plans on AMR – A review and country experiences