prepared by Rémi Gschwind
This February issue presents studies on the presence of antibiotic resistance genes, as usual, in various environments according to the One Health concept. Human microbiome is considered as a reservoir of antibiotic resistance genes and in this digest we have a special highlight on the effect of antibiotics on newborns microbiome. Hospital environment is also discussed with methods for prediction of resistance and resistance prevalence in several context. Finally, studies focusing on antibiotic resistance in a more environmental perspective are also presented to complete this One Health circle.
Human microbiome
Impact of long-term dietary habits on the human gut resistome in the Dutch population – Paul B. Stege – Scientific Reports
Effects of early-life antibiotics on the developing infant gut microbiome and resistome: a randomized trial – Marta Reyman – Nature Communications
In addition to the undeniable beneficial effects of antibiotics on newborns life span, antibiotics also have deleterious side effects such as microbiome species richness depletion and antibiotic resistant bacteria selection. Side effects could be even more deleterious if we take into account side effects that might have detrimental effects later in life according to the developmental origin of health and disease concept. Reyman et al. conducted the ZEBRA study enrolling 147 infants born at term either by natural delivery or cesarean section, for whom broad spectrum antibiotics were used. Reduced gut microbial diversity and prolonged ecological perturbations were detected compared with healthy term-born controls (still measurable after 12 months). Also, shifts in AMR gene profile were evidenced using qPCR and confirmed by metagenomic shotgun sequencing of a subset of samples. Those effects were different depending on the antimicrobial use (penicillin+gentamicin being the least deleterious) highlighting antibiotics choice importance.
Gut microbiome signatures and host colonization with multidrug-resistant bacteria – Nicole S.Isles – Trends in Microbiology
Hospital environment
Extended-Spectrum Beta-Lactamase and Carbapenemase-Producing prediction in Klebsiella pneumoniae based on MALDI-TOF mass spectra – Alejandro Guerrero-López – bioRxiv
Genomic diversity and antimicrobial resistance of Prevotella species isolated from chronic lung disease airways – Kasey A. Webb –Microbial Genomics
Long read sequencing reveals genomic diversity and associated plasmid movement of carbapenemase-producing bacteria in a UK hospital over six years – Leah Roberts – OSF Preprints
Environmental antibiotic resistance
Deciphering the extracellular and intracellular antibiotic resistance genes in multiple environments reveals the persistence of extracellular ones – Yina Zou – Journal of Hazardous Materials
Long-read metagenomic sequencing reveals shifts in associations of antibiotic resistance genes with mobile genetic elements from sewage to activated sludge – Dongjuan Dai – Microbiome
Carriage of antibiotic resistant bacteria in endangered and declining Australian pinniped pups – Mariel Fulham – PlosOne
A new insight into the ARG association with antibiotics and non-antibiotic agents—antibiotic resistance and toxicity – Shaojing Sun – Environmental Pollution
Impact of fertilization with pig or calf slurry on antibiotic residues and resistance genes in the soil – Huygens Judith – Science of The Total Environment
Monitoring and evaluation of antibiotic resistance genes in three rivers in northeast China – Chen Zao – Environmental Science and Pollution Research
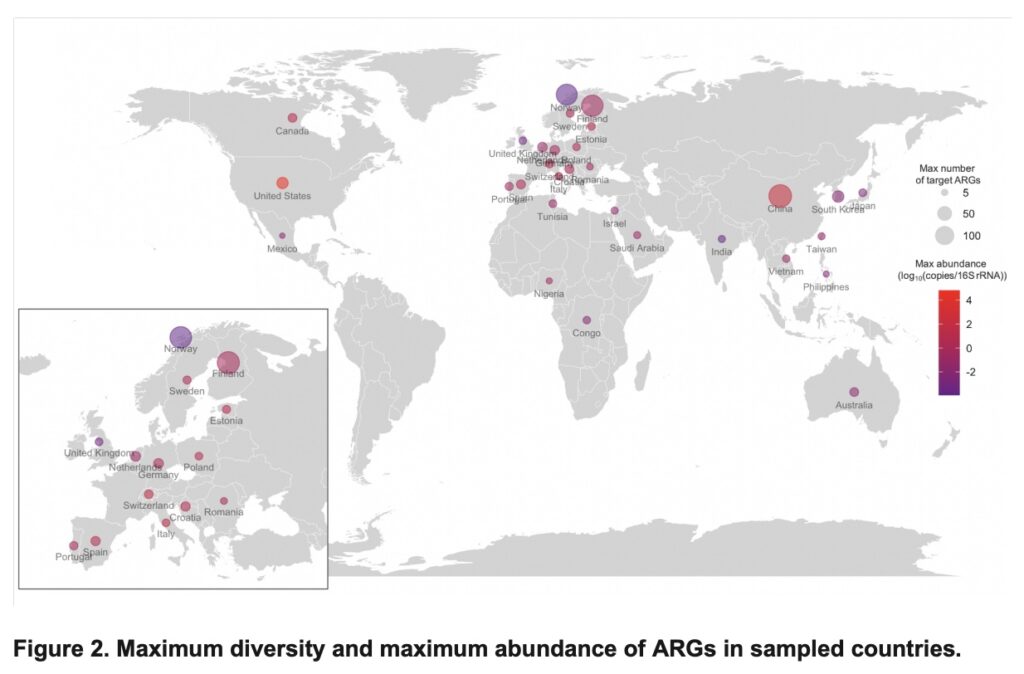
Meta-analysis reveals the global picture of antibiotic resistance gene prevalence across environments – Anna Abramova – bioRxiv
Fundamental microbiology
Evolution of ColE1-like plasmids across γ-Proteobacteria: From bacteriocin production to antimicrobial resistance – Manuel Ares-Arroyo – PLOS Genetics
Tolerance and resistance of microbial biofilms – Oana Ciofu – Nature Reviews – Microbiology
Gradients in gene essentiality reshape antibacterial research – Andrew M Hogan – FEMS Microbiology Reviews
The evolution of colistin resistance increases bacterial resistance to host antimicrobial peptides and virulence – Pramod K. Jangir – bioRxiv
Colistin is an efficient antimicrobial peptide (AMP) used at a large scale in agriculture in the 1980s. Today, it is being used as “last-resort” antimicrobial to treat infections. One serious concern lies in the potential cross resistance between colistin resistance and host AMP resistance (since they have common physicochemical properties and mechanisms) which could increase pathogen transmission and virulence. Colistin resistance is mainly due to MCR-1 which became widely distributed across all niches because of bacterial migration and horizontal transfer. Here, Jangir et al. tested the hypothesis that evolving colistin resistance via MCR genes acquisition promotes resistance in bacteria against host AMPs. The presence of MCR indeed increased resistance against several AMPs coming from different sources. It highlights the importance of assessing the impact of evolved resistance to future therapeutic AMPs.
Courses
Emerging laboratory and point-of-care technologies for detection of AMR and bacterial infection in veterinary medicine – ESCMID, 9 March 2022
Workshops, Seminars &c
Risk assessment of biocide and antibiotic resistance – BIOCIDE consortium, 9 March 2022
Plasmids as vehicles of AMR spread – The Abdus Salam International Centre for Theoretical Physics, 21-25 March 2022
CHARM Virtual Seminar Series 2022, Bi-weekly Tuesdays 9:00-10-00 am