prepared by Marcus Wenne
In September, the EMBARK team got together to:
1. discuss current and future AMR research efforts and
2. have fun!!
If you live in the Northern Hemisphere every day is becoming darker and colder when we are steadily approaching winter. It is therefore extra important to illuminate your mind with interesting research and get that warm feeling in your body when reading a well written and structured paper. This is hopefully the feeling you will get when reading our October AMR digest. It touches the topics of evolution, the release of new databases, biofilms, wastewater and so much more. If you are from the Southern Hemisphere, you are of course equally welcome to take part of this interesting collection of papers!
Wastewater
Predicting selection for antimicrobial resistance in UK wastewater and aquatic environments: Ciprofloxacin poses a significant risk – April Hayes – Environment International
Microbiome and resistome dynamics along a sewage-effluent-reservoir continuum underline the role of natural attenuation in effluent receiving reservoirs – Inês Leão, Leron Khalifa, Nicolas Gallois, Ivone Vaz-Moreira, Uli Klümper, Daniel Youdkes, Shaked Palmony, Lotan Dagai, Thomas U. Berendonk, Christophe Merlin, Célia M. Manaia, Eddie Cytryn – bioRxiv
New methods, tools or approaches
Evaluation of FEAST for metagenomics-based source tracking of antibiotic resistance genes – Jinping Chen – Journal of Hazardous Materials
Determination and quantification of microbial communities and antimicrobial resistance on food through host DNA-depleted metagenomics – Samuel J Bloomfield – Food Microbiology
Quantification of the mobility potential of antibiotic resistance genes through multiplexed ddPCR linkage analysis – Magali de la Cruz Barron, David Kneis, Alan Xavier Elena, Kenyum Bagra, Thomas U. Berendonk, Uli Klümper – bioRxiv
Biofilm
Metagenomic insights into taxonomic, functional diversity and inhibitors of microbial biofilms – Madangchanok Imchen – Microbiological Research
Biofilm antimicrobial susceptibility through an experimental evolutionary lens – Tom Coenye – Biofilms and Microbes
Databases & resources
*ResFinderFG v2.0: a database of antibiotic resistance genes obtained by functional metagenomics – Rémi Gschwind – bioRxiv
A bioinformatic analysis is not better than the database and tools you base it on. Here Rémi
Gschwind et al. describes a new and updated version of ResFinderFG, version 2.0. According
to the authors most genes in antibiotic resistance gene databases mostly originate from
culturable and pathogenic bacteria. ResfinderFG 2.0 is instead a database based on a
literature search on studies identifying resistance genes using functional metagenomics.
This means that there is a reduced bias in the ResFinderFG v.2.0 database towards
culturable pathogens.
Ab-AMR: A Comprehensive Repository of Acinetobacter baumannii to Understand the Molecular Landscape of Antimicrobial Resistance – Tina Sharma – bioRxiv
CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database – Brian P Alcock – Nucleic Acids Research
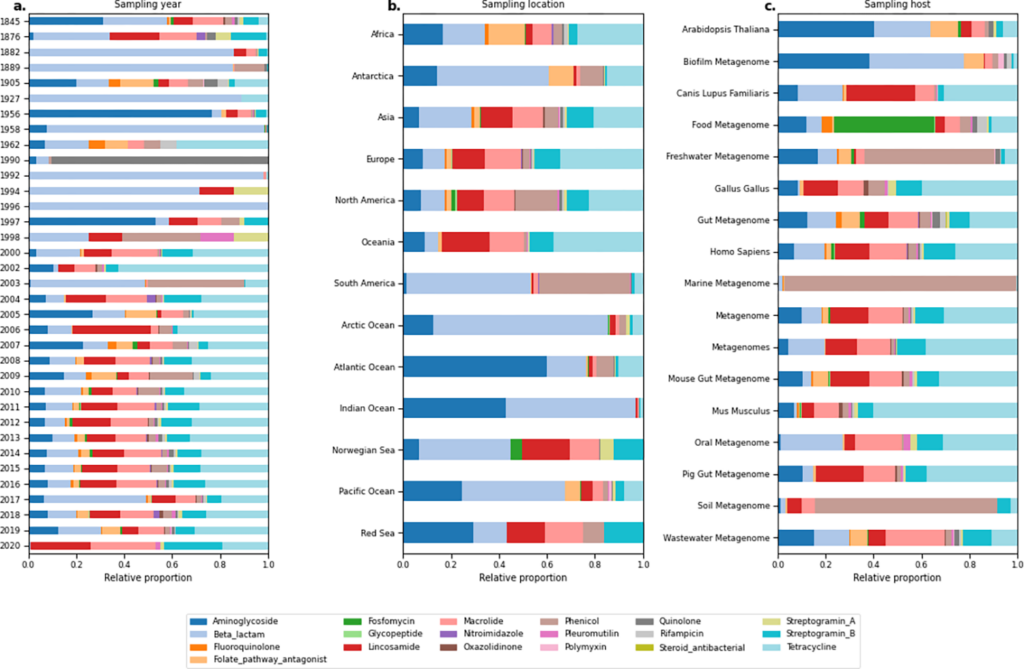
*A curated data resource of 214K metagenomes for characterization of the global antimicrobial resistome – Hannah-Marie Martiny – PLOS BIOLOGY
Large scale metagenomic screenings of the worlds entire catalogue of metagenomic data
could potentially reveal interesting global and local patterns of antibiotic resistance.
Undertaking such a task is however way to computer intensive for most research groups to
undertake. Luckily, Hannah-Marie Martiny et al. have already undertaken this task. In this
paper they present a publicly available resource of antibiotic resistance gene abundance
based on 442 Tbp of sequencing reads from 214,095 metagenomic samples from the
European Nucleotide Archive.
Reviews
Time for Some Group Therapy: Update on Identification, Antimicrobial Resistance, Taxonomy, and Clinical Significance of the Bacteroides fragilis Group – Sophonie Jean – Journal of Clinical Microbiology
Tackling AMR from a multidisciplinary perspective: a primer from education and psychology – Alicia Calvo‑Villamañán – International Microbiology
Machine learning in predicting antimicrobial resistance: a systematic review and meta-analysis – Rui Tang – International Journal of Antimicrobial Agents
A bottom-up view of antimicrobial resistance transmission in developing countries – Odion O. Ikhimiukor – Nature Microbiology
Plasmids
Phage-Plasmids Spread Antibiotic Resistance Genes through Infection and Lysogenic Conversion – Eugen Pfeifer – mBio
New antibiotics
Small Molecule Antibiotics Inhibit Distinct Stages of Bacterial Outer Membrane Protein Assembly – Janine H Peterson – mBio
Ring-fused 2-pyridones effective against multidrug-resistant Gram-positive pathogens and synergistic with standard-of-care antibiotics – Taylor M Nye – PNAS
Clinical studies
Oropharyngeal and intestinal concentrations of opportunistic pathogens are independently associated with death of SARS‑CoV‑2 critically ill adults – Juliette Patrier, Khanh Villageois‑Tran, Piotr Szychowiak, Stéphane Ruckly, Rémi Gschwind, Paul‑Henri Wicky, Signara Gueye, Laurence Armand‑Lefevre, Mehdi Marzouk, Romain Sonneville, Lila Bouadma, Marie Petitjean, Fariza Lamara, Etienne de Montmollin, Jean‑Francois Timsit, Etienne Ruppé and The French COVID Cohort Study Group – Critical Care
Genomic characterisation of multidrug-resistant Escherichia coli, Klebsiella pneumoniae, and Acinetobacter baumannii in two intensive care units in Hanoi, Viet Nam: a prospective observational cohort study – Leah W Roberts – Lancet Microbe
Evolution
*The evolution of spectrum in antibiotics and bacteriocins – Jacob D Palmer and Kevin R
Foster – PNAS
An important weapon in microbial competition is the production of antibiotics. But when is
it preferential for a microbial population to produce wide or narrow spectrum antibiotics?
In this paper Jacob D Palmer and Kevin R Foster have used an evolutionary model approach
in an attempt to shed light on this question.
Antibiotics in the environment
Deciphering chloramphenicol biotransformation mechanisms and microbial interactions via integrated multi‑omics and cultivation‑dependent approaches – Jiayu Zhang, Xiaoyan Li, Uli Klümper, Huaxin Lei, Thomas U. Berendonk, Fangliang Guo, Ke Yu, Chao Yang & Bing Li – Microbiome
Antibiotic resistance mechanisms
L-Form Switching in Escherichia coli as a Common b-Lactam Resistance Mechanism – Aleksandra P Fabijan – Microbiology Spectrum
Antibiotic usage
Factors influencing usage of antimicrobial drugs among pastoralists in Kenya – Dennis N Makau – Tropical Animal Health and Production
Soil
Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics – Hong-Zhe Li – PNAS
Anthropogenic impact
Leave No Trace? Ecological and anthropogenic determinants of antibiotic resistant bacteria in a recreational alpine environment – Laura C Scott – Environmental Research
Podcasts
Microbiology Lab Pod September 2022 Pod: Environmental Antibiotic Resistance – Johan Bengtsson-Palme lab
Webinars
Vaccination as a strategy to combat antimicrobial resistance – Organised by the (CSA) DESIGN One Health AMR, together with MRC, BactiVac